Joseph Krzycki

Contact Information
Professor of Microbiology
Areas of Expertise
- Biochemistry and molecular biology of methanogenic archaea
Education
- B.S. University of Nebraska, Lincoln
- M.S. University of Wisconsin, Madison
- Ph.D. University of Wisconsin, Madison
Research Interests
Methanogenesis and Pyrrolysine, the 22ndAmino Acid
Our lab is interested in the biochemistry of biological methane production by methanogenic Archaea. People are drawn to research on methanogens for a number of reasons. These microbes inhabit anaerobic environments like freshwater and marine sediments or municipal waste digesters. In these environments, methane formation is a critical final step in the degradation of organic matter. Methanogenesis produces cellular energy even while producing a human energy source. Methanogens are the charter members of the Archaea, a Domain of life evolutionarily separate from Bacteria or Eucarya. Befitting this unique evolutionary status, methanogens possess an unusual biochemistry and molecular biology, and have evolved distinctive cofactors and enzymes to form methane. In recent years we have shown that some methanogens have even changed their genetic code to suit the needs of methanogenesis. To use methylamines to make methane, they synthesize proteins possessing an unusual amino acid call pyrrolysine. Pyrrolysine is inserted during translation, making it the 22ndamino acid found to have entered the genetic code. While we first found pyrrolysine in methanogens, evidence for pyrrolysine has now been found in nine different genera of Bacteria and Archaea, including a human gut bacterium.
Methanogenesis from Methylamines
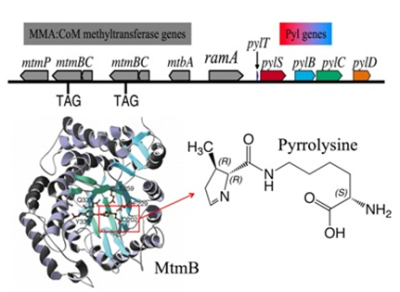
Our lab began studyingMethanosarcinaspp. because this genus has evolved an expanded range of substrates that includes, among other substrates, methylamines such as trimethylamine, dimethylamine, and monomethylamine (MMA). These methylamines are produced from the breakdown of lipid components or cell osmolytes and are abundantly produced in some fresh and marine waters. We used classical biochemical techniques to identify proteins converting the three methylamines to methyl-Coenzyme M (CoM). Methyl-CoM is the precursor of methane, carbon dioxide, and much of cellular carbon. Three types of methylamine methyltransferases use specifically either trimethylamine, dimethylamine, or monomethylamine to methylate a partner corrinoid protein, which are then substrates of a fourth methyltransferase which forms methyl-CoM. As we characterized the proteins and sequenced their genes, we were surprised to find in-frame amber (TAG) codons interrupting the genes encoding each type of methylamine methyltransferase. We established that the amber codons are translated inMethanosarcinaspp. Our lab then collaborated with Michael Chan, who solved the structure of MtmB, the MMA methyltransferase, and deduced the structure of the new amino acid corresponding to the amber codon. We named the new amino acid pyrrolysine.
The Genetic Encoding of Pyrrolysine
Our subsequent studies have shown that pyrrolysine directly entered the genetic code much like the original twenty amino acids. Pyrrolysine has its own tRNA (called tRNAPylor tRNACUA) that can decode UAG during translation. A gene calledpylSfollows the tRNAPylgene in the methanogen genome. The protein made frompylSis a pyrrolysyl-tRNA synthetase. That is, this enzyme attaches chemically synthesized pyrrolysine directly to tRNAPyl, a key step that allows pyrrolysine entrance to the genetic code. We demonstrated that methanogenpylSandpylTfunction not only in vitro, but also in vivo inE. coli. This set the stage for later exploitation of PylS and tRNAPylto insert useful chemically modifiable residues into recombinant proteins, a strategy now pursued by a number of labs.We have found the methanogen still uses UAG as a stop codon; and a balance exists between use of UAG as a stop codon and a sense codon withinMethanosarcina acetivorans. Downstream sequences can enhance UAG read-through in at least one methylamine methyltransferase transcript, but amber codons are translated in foreign reporter genes as well, probably via amber suppression by tRNApyl.
Biosynthesis of Pyrrolysine
Nearby the pyrrolysyl-tRNA synthetase and tRNAPylgenes are three morepylgenes,pylB,pylC, andpylD. The enzymes from these genes are responsible for making pyrrolysine from common cellular metabolites. We have shown that transfer of the these three genes, along withpylTandpylS, allows a common bacteria likeE. colito make pyrrolysine and insert it into protein under the direction of UAG codons. In other words, these five genes program the bacteria to make pyrrolysine and incorporate it into its genetic code. We recently used this recombinant system to elucidate pyrrolysine biosynthesis. We have shown pyrrolysine is made entirely from lysine, and have proposed a pathway in which PylB converts lysine to a methylated ornithine derivative, which is joined to another lysine by PylC, before oxidation by PylD results in the formation of pyrrolysine. This work has set the stage for detailed in vitro analysis of the enzymes. These results also have implications for both understanding the evolution of the genetic code, as well as for the practical applications of pyrrolysine derivatives in production of recombinant proteins with chemically modifiable residues.
Function of Pyrrolysine in Methylamine Methyltransferases
Deletion of the tRNAPylgene fromMethanosarcina acetivoransresults in an inability to make methane from methylamines, but no other detectable defects in the organism. This suggests pyrrolysine is necessary for methylamine methyltransferase synthesis, but why is it present in the three types of methylamine methyltransferases, proteins that share no sequence similarity? The answer will help explain how methane is made from methylamines, but may also provide a rationale for the fact that pyrrolysine found entrance into the genetic code. The most favored current hypothesis is that pyrrolysine forms a methylammonium adduct as a preliminary to demethylation by the corrinoid protein, but other scenarios are possible. We have recently shown that replacement of pyrrolysine with other amino acids severely compromises the activity of the protein, but muchin vitrowork remains to understand the role of pyrrolysine in the methylamine methyltransferases.
RamA, a Novel iron-sulfur Protein that Activates the Methylamine Methyltransferases.
Another facet of the methylamine methyltransferases is how these corrinoid dependent methyltransferase reactions are “activated.” We have found that a single iron-sulfur protein, RamA, carries out the ATP dependent reduction of the cobalt ion of methylamine corrinoid proteins to the Co(I) state, thereby activating them as a substrate for their partner methylamine methyltransferase. RamA is a member of a large group of iron-sulfur proteins whose function was previously unknown, and these proteins are very unlike other iron-sulfur proteins that accomplish analogous difficult reductions. How this novel protein family ties ATP hydrolysis to the reduction of the corrinoid protein is an important question for our future research.
Memberships
Relevant Publications
Gaston MA,Zhang L,Green-Church KB&Krzycki JA. (2011). The complete biosynthesis of the genetically encoded amino acid pyrrolysine from lysine. Nature 471:647-650.
Ferguson DJ,Longstaff DG &Krzycki JA. (2011). Assay of methylotrophic methyltransferases from methanogenic Archaea. Methods in Enzymology: 494:139-158.
Gaston MA, Jiang R & Krzycki JA. (2011) Functional context, biosynthesis, and genetic encoding of pyrrolysine. Current Opinions in Microbiology, in press (invited review).
Rother M &Krzycki JA. (2010) Selenocysteine, pyrrolysine, and the unique energy metabolism of methanogenic archaea. Archaea (an open access journal) pii: 453642; doi:10.1155/2010/453642.
Ferguson T, Soares JA, Lienard T, Gottschalk G &Krzycki JA. (2009) RamA, a protein required for reductive activation of corrinoid-dependent methylamine methyltransferase reactions in methanogenic archaea. J. Biol. Chem. 284(4):2285-2295.
Li WT, Mahapatra A, Longstaff DG, Bechtel J, Zhao G, Kang PT, Chan MK& Krzycki JA. (2009) Specificity of pyrrolysyl-tRNA synthetase for pyrrolysine and pyrrolysine analogs. J. Mol. Biol.385(4):1156-64.
Longstaff DG, Larue RC, Faust JE, Mahapatra A, Zhang L, Green-Church KB &Krzycki JA. (2007) A natural genetic code expansion cassette enables transmissible biosynthesis and genetic encoding of pyrrolysine. Proc. Natl. Acad. Sci. U S A. 104(3):1021-1026.
Mahapatra A,Patel A,Soares JA,Larue RC,Zhang JK,Metcalf WW &Krzycki JA. (2006) Characterization of a Methanosarcina acetivorans mutant unable to translate UAG as pyrrolysine. Mol. Microbiol. 59:56-66.
Soares JA,Zhang L,Pitsch RL,Kleinholz NM,Jones RB,Wolff JJ,Amster J,Green-Church KB &Krzycki JA. (2005) The residue mass of L-pyrrolysine in three distinct methylamine methyltransferases. J. Biol. Chem. 280:36962-36969.
Blight SK,Larue RC,Mahapatra A,Longstaff DG,Chang E,Zhao G,Kang PT,Green-Church KB,Chan MK&Krzycki JA. (2004) Direct charging of tRNACUA with pyrrolysine in vitro and in vivo. Nature. 431:333-335.
Srinivasan G,James CM &Krzycki JA. (2002) Pyrrolysine encoded by UAG in Archaea:charging of a UAG-decoding specialized tRNA. Science. 296:1459-1462.
Hao B,Gong W,Ferguson TK,James CM,Krzycki JA &Chan MK.(2002) A new UAG encoded residue in the structure of a methanogen methyltransferase. Science. 296: 1462-1465.
Paul L, Ferguson Jr DJ& Krzycki JA. (2000) The trimethylamine methyltransferase gene and multiple dimethylamine methyltransferase genes of Methanosarcina barkeri contain in-frame and read-through amber codons. J Bacteriol. 182:2520-2529.
Two Seminal Publications
Burke SA,Lo SL &Krzycki JA.(1998) The clustered genes encoding the methyltransferases of methanogenesis from monomethylamine. J. Bacteriol. 180:3432-3440. (The first description of an amber codon in a methylamine methyltransferase and the hypothesis that it might encode an unusual amino acid).
Burke SA&Krzycki JA(1997) Reconstitution of monomethylamine:coenzyme M methyl transfer with a corrinoid protein and two methyltransferases purified from Methanosarcina barkeri. J. Biol. Chem. 272:16570-16577. (The first isolated methylamine:CoM methyltransferase system functional in vitro).
