Tina Henkin

Contact Information
Professor of Microbiology Emeritus; Previously: Distinguished University Professor, Robert W. and Estelle S. Bingham Professor of Biological Sciences
Areas of Expertise
- Transcription termination control in Gram-positive bacteria
- RNA structure/function
- Riboswitches
Education
- B.S., Swarthmore College, 1977
- Ph.D., University of Wisconsin-Madison, 1984
- Postdoc, Tufts University School of Medicine, 1984-1987
Affiliations
- Center for RNA Biology
- Molecular, Cellular, and Developmental Biology Graduate Program
- The Ohio State Biochemistry Program
Awards
- Distinguished University Professor, Ohio State University, 2016
- Member, National Academy of Sciences, 2012
- Fellow, American Academy of Arts & Sciences, 2009
- Robert W. and Estelle S. Bingham Professor of Biological Sciences, 2009
- National Academies of Science Pfizer Prize in Molecular Biology, 2006
- Fellow, American Association for the Advancement of Science, 2005-present
- Chair, American Society for Microbiology, Division H, 2005-2006
- Distinguished Scholar Award, Ohio State University, 2004
- Fellow, American Academy of Microbiology, 2003
Research Interests
The main area of interest in our laboratory is the analysis of the mechanisms through which cells sense changes in their environment and transmit that information to the level of gene expression.We use the Gram-positive bacterium Bacillus subtilis as a model system, and we focus primarily on genes involved in protein synthesis and amino acid metabolism.We have uncovered systems in which nascent RNA transcripts act as riboswitches to directly sense physiological signals and control gene expression through RNA structural rearrangements.
Nascent RNAs can sense uncharged tRNA: the T box riboswitch
Characterization of the B. subtilis tyrS gene, encoding tyrosyl-tRNA synthetase, revealed a novel mechanism of gene regulation at the level of transcription antitermination. The tyrSgene is a member of a large family of aminoacyl-tRNA synthetase and amino acid biosynthesis genes in Gram-positive bacteria that are regulated by a common mechanism.Each gene in this family responds individually to limitation for the appropriate amino acid. Amino acid limitation is monitored via interaction of the 5’ region of the nascent transcript with the cognate uncharged tRNA.This interaction is directed by pairing of the anticodon of the tRNA with a single codon, designated the "Specifier Sequence," in the mRNA.The mRNA-tRNA interaction occurs in the absence of translation, and antitermination can occur in a purified transcription system with no additional cellular factors, indicating that the mRNA is sufficient for specific recognition of the cognate tRNA.We are currently investigating the molecular details of the leader RNA-tRNA interaction, and the structural shifts in both RNA partners that occur upon binding.We are also testing novel antibiotics for their ability to target the T box mechanism.
Nascent RNAs can sense small molecules: metabolite-binding riboswitch RNAs
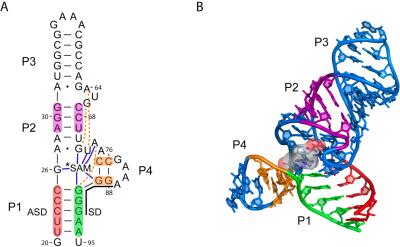
Analysis of genes involved in methionine metabolism revealed a second global transcription antitermination system, dedicated to genes in this pathway.Like the T box system, the S box system is widely used in Gram-positive organisms. Genes regulated by this mechanism contain highly conserved sequence and structural elements in their mRNAs, and expression is induced by starvation for methionine. We have now shown that the molecular effector for this system is S–adenosylmethionine (SAM), which binds directly to the leader RNA and modulates its structure to promote transcription termination. A second SAM-binding RNA, the SMK box, was identified in lactic acid bacteria and shown to regulate gene expression at the level of translation initiation.We have also shown that lysine biosynthesis genes are regulated by a similar mechanism, with specific leader RNA binding of lysine.Current work is focusing on the molecular mechanisms of effector recognition and RNA rearrangement in response to effector binding.
Selected Publications
Textbook: Henkin, T. M., and J. E. Peters. 2020. Snyder & Champness Molecular Genetics of Bacteria, 5th Edition. American Society for Microbiology/Wiley, Washington, DC.
Bhagdikar, D., F. J. Grundy and T. M. Henkin. 2020. Transcriptional and translational S-box riboswitches differ in ligand-binding properties. J. Biol. Chem. 295: 6849-6860.
Kreuzer, K. D. and T. M. Henkin. 2018. The T box riboswitch: tRNA as an effector to modulate gene regulation. Microbiol Spectr. 6(4). doi: 10.1128/microbiolspec.RWR-0028-2018.
Sherwood, A. V., J. V. Frandsen, F. J. Grundy and T. M. Henkin. 2018. New tRNA contacts facilitate ligand binding in a Mycobacterium smegmatis T box riboswitch. Proc Natl. Acad. Sci. USA 115: 3894-3899.
Sherwood, A.V. and T. M. Henkin. 2016. Riboswitch-mediated gene regulation: Novel RNA architectures dictate gene expression responses. Annu. Rev. Microbiol. 70: 361-374.
Sherwood, A. V., F. J. Grundy and T. M. Henkin. 2015. T box riboswitches in Actinobacteria: translational regulation via novel tRNA interactions. Proc. Natl. Acad. Sci. USA 112: 1113-1118.
Grigg, J. C., Y. Chen, F. J. Grundy, T. M. Henkin, L. Pollack and A. Ke. 2013. T box RNA decodes both the information content and geometry of tRNA to affect gene expression. Proc. Natl. Acad. Sci. USA. 110: 7240-7245.
Wilson, R. C., A. M. Smith, R. T. Fuchs, I. R. Kleckner, T. M. Henkin, and M. P. Foster. 2011. Tuning riboswitch regulation through conformational selection. J. Mol. Biol. 405: 926-938.
Smith, A. M., R. T. Fuchs, F. J. Grundy and T. M. Henkin. 2010. The SMK box of Enterococcus faecalis is a reversible riboswitch. Mol. Microbiol. 78: 1393-1402.
Lu, C., F. Ding, A. Chowdhury, V. Pradhan, J. Tomsic, M. W. Holmes, T. M. Henkin and A. Ke. 2010. SAM recognition and conformational switching mechanism in the Bacillus subtilis yitJ S Box/SAM-I riboswitch. J. Mol. Biol. 404: 803-818.
Grundy, F. J. and T. M. Henkin. 1993. tRNA as a positive regulator of transcription antitermination in B. subtilis. Cell 74: 475-482.
PubMed
